Antibody
Engineering
Home » Antibody Engineering
Improve antibodies for your
drug development pipeline
Our integrated YUMAB® platform enables not only fast-track discovery but also engineering of existing antibodies. The art of antibody engineering involves a combination of in-vitro and AI-assisted in silico technologies to optimize clinically relevant properties, such as affinity, stability, specificity, and selectivity.
As a special engineering service, YUMAB offers AI-assisted antibody humanization.

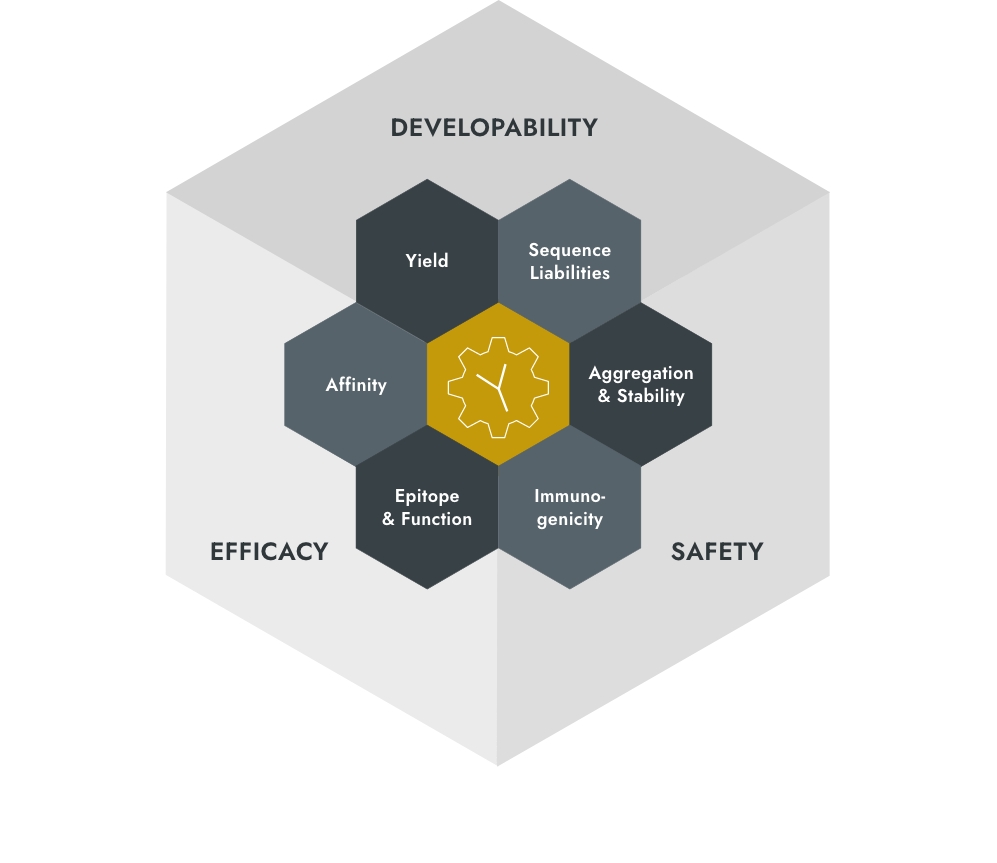
Paying attention to developability,
safety and efficacy
Our main objectives:
- Generate a broad diversity pool for selection
- Remove sequence liabilities
- Enable new functions
- Improve affinities
- Achieve high stability
- Guarantee low immunogenicity
Our engineering principle:
In vitro evolution – zooming in on the right fit
At YUMAB, we use AI-assisted rational mutagenesis to increase the antibody diversity combined with in-vitro evolution technologies, for the selection of antibodies with improved features or functional variations.
In silico analysis – de-risking sequence check
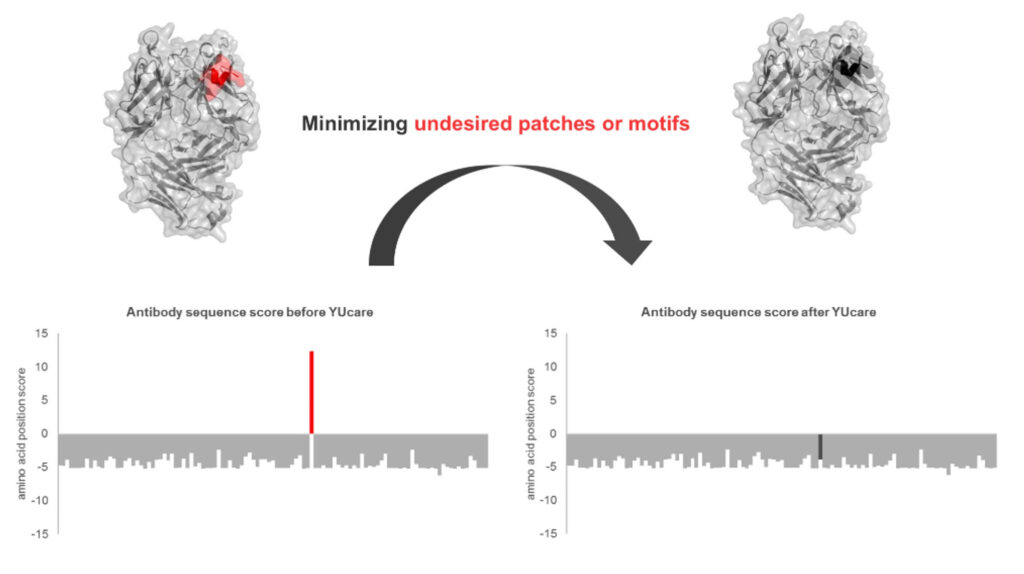
YUcare is YUMAB’s AI-assisted in silico sequence optimization tool, that we use to model antibody developability characteristics. We also compare the sequences to late-stage and market-approved antibodies and remove unwanted patches or motifs from the top candidates. This approach is not only time- and cost-saving but also guarantees a high success rate.
We then produce and test the selection of ‘polished’ antibodies in their final format to confirm functionality and manufacturability.
Taking care of developability with YUcare
YUMAB’s proprietary YUcare toolbox is a suite of in silico tools that supports lead candidate selection and enhances lead development. The developability service provides the analysis of biochemical and biophysical properties of antibody lead candidates and is used to remove liabilities in the antibody sequence and structure as well as to accelerate antibody drug development.
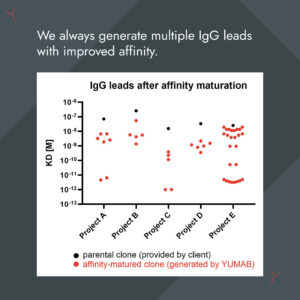
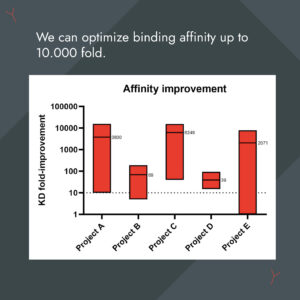
We're experts in affinity optimization


YUMAB Case Study
YUMAB collaboration with Merck KGaA
Bispecific PROxAb antibody shuttle
against difficult targets
The selective degradation of disease-related proteins by PROTACs (Proteolysis-targeting chimeras ) is one approach that is currently explored in a number of drug development programs. Schneider (Merck KGaA) and colleagues have introduced PROxAb shuttles as next generation PROTACs using antibodies in a modular plug-and-play platform. In collaboration with YUMAB the group developed PROxAb, a PROTAC specific antibody fused to a therapeutic antibody thereby creating a bispecific antibody for tumor therapy. YUMAB has extensive expertise in the development of camelid derived single domain (VHH) antibodies which, due to their small size and elongated CDR3 loop, can interact with difficult to access epitopes. PROxAbs enable the development of new complementary therapies for many difficult to access targets in a broad range of diseases.